Status: Done started: 2018-Jan-15 14:26:58 UTC
| Project Name | C10F1 |
| Project Name | C10F1 |
| Cluster # | 1 | 2 | 3 | 4 | 5 | 6 | 7 | 8 | 9 | 10 | 11 | 12 |
| Cluster density | 104.6 | 102.0 | 92.0 | 85.1 | 80.8 | 77.5 | 70.0 | 67.6 | 58.9 | 58.2 | 55.2 | 52.6 |
| Cluster size | 220 | 229 | 194 | 191 | 182 | 169 | 166 | 157 | 132 | 126 | 123 | 111 |
| Average cluster RMSD | 2.1 | 2.2 | 2.1 | 2.2 | 2.3 | 2.2 | 2.4 | 2.3 | 2.2 | 2.2 | 2.2 | 2.1 |
| # | 1 | 2 | 3 | 4 | 5 | 6 | 7 | 8 | 9 | 10 | 11 | 12 |
| RMSD | 4.07 | 4.51 | 4.33 | 3.85 | 3.48 | 3.82 | 4.18 | 3.92 | 4.32 | 3.89 | 3.78 | 4.12 |
| GDT_TS | 0.53 | 0.51 | 0.51 | 0.53 | 0.57 | 0.57 | 0.51 | 0.54 | 0.56 | 0.56 | 0.57 | 0.52 |
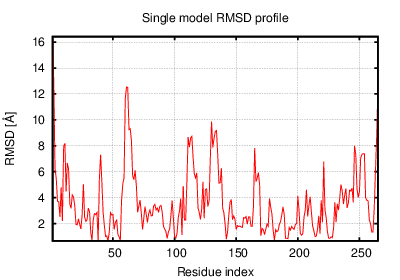
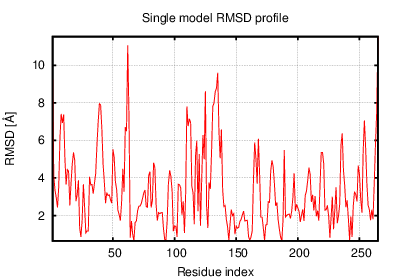
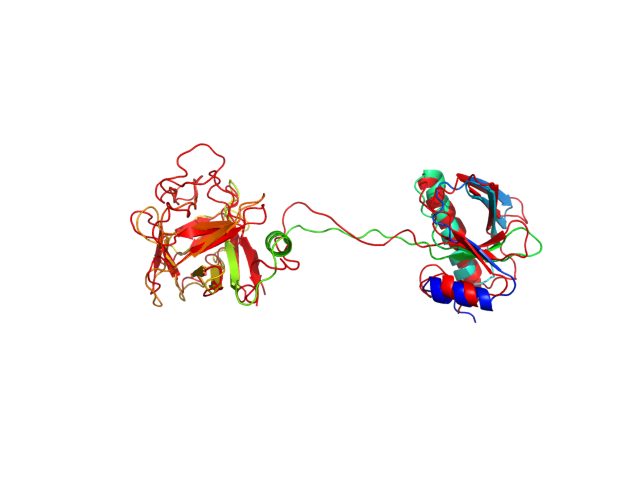
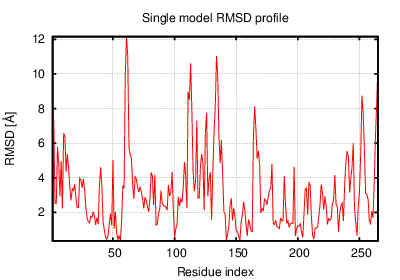
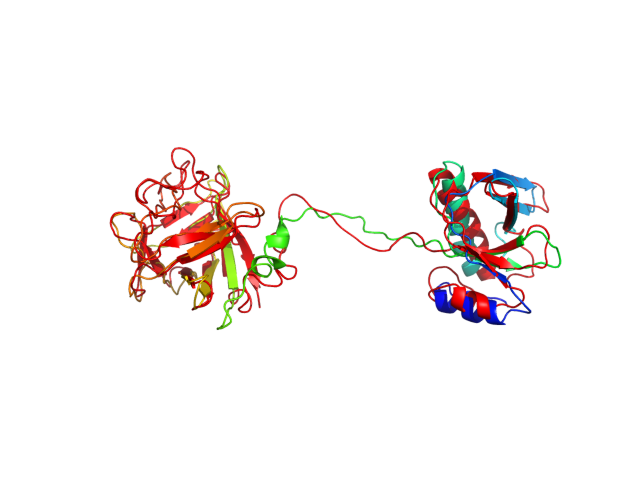
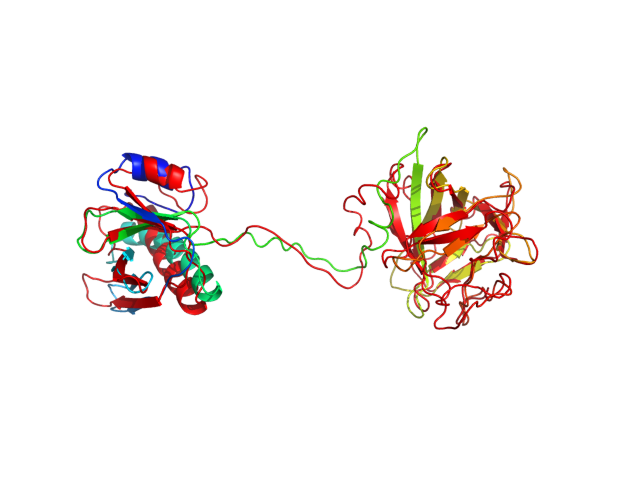
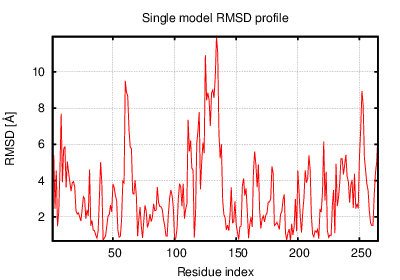
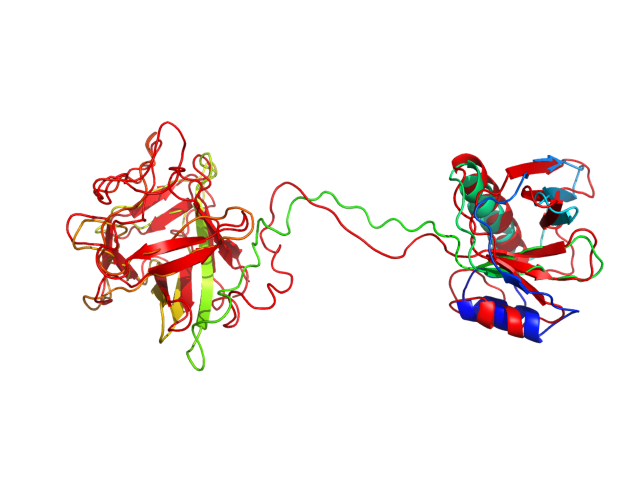
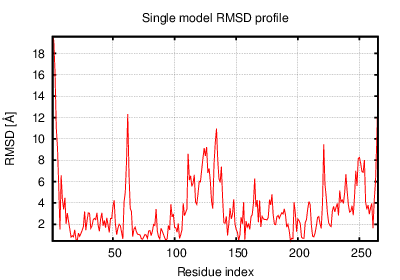
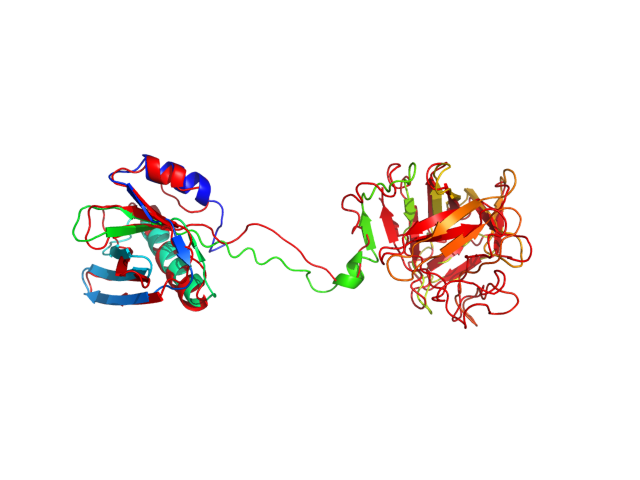
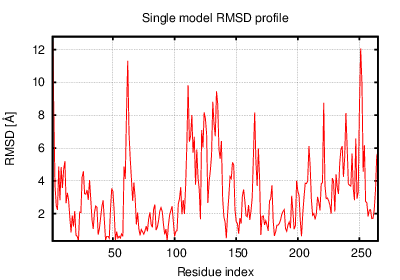
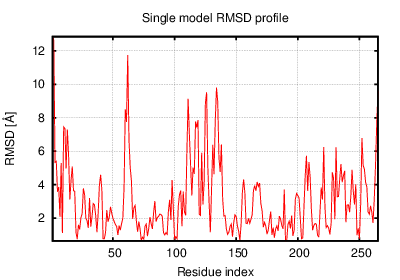
The table contains RMSD and GDT_TS values (calculated on the Cα atoms) between the predicted models and the input structure. Note that GDT_TS metric is intended as a more accurate measurement than the more common RMSD.
Read more about the root-mean-square deviation (RMSD) measure
Read more about the global distance test (GDT, also written as GDT_TS to represent "total score") measure.
| # | 1 | 2 | 3 | 4 | 5 | 6 | 7 | 8 | 9 | 10 | 11 | 12 |
| 1 | 0.00 | 3.34 | 3.59 | 4.45 | 3.68 | 3.03 | 3.58 | 3.57 | 3.41 | 2.85 | 3.11 | 3.28 |
| 2 | 3.34 | 0.00 | 4.19 | 4.91 | 3.33 | 3.01 | 3.89 | 4.13 | 4.32 | 3.74 | 4.00 | 3.55 |
| 3 | 3.59 | 4.19 | 0.00 | 4.05 | 4.17 | 3.59 | 3.46 | 3.20 | 4.11 | 3.22 | 2.87 | 3.81 |
| 4 | 4.45 | 4.91 | 4.05 | 0.00 | 3.76 | 4.05 | 3.58 | 3.56 | 4.81 | 4.01 | 3.54 | 4.72 |
| 5 | 3.68 | 3.33 | 4.17 | 3.76 | 0.00 | 2.81 | 3.60 | 3.44 | 4.48 | 3.67 | 3.59 | 3.34 |
| 6 | 3.03 | 3.01 | 3.59 | 4.05 | 2.81 | 0.00 | 3.40 | 3.11 | 4.24 | 3.34 | 3.22 | 3.22 |
| 7 | 3.58 | 3.89 | 3.46 | 3.58 | 3.60 | 3.40 | 0.00 | 3.02 | 4.50 | 3.44 | 2.91 | 3.84 |
| 8 | 3.57 | 4.13 | 3.20 | 3.56 | 3.44 | 3.11 | 3.02 | 0.00 | 4.33 | 3.21 | 2.63 | 3.67 |
| 9 | 3.41 | 4.32 | 4.11 | 4.81 | 4.48 | 4.24 | 4.50 | 4.33 | 0.00 | 3.79 | 3.89 | 3.97 |
| 10 | 2.85 | 3.74 | 3.22 | 4.01 | 3.67 | 3.34 | 3.44 | 3.21 | 3.79 | 0.00 | 2.69 | 3.38 |
| 11 | 3.11 | 4.00 | 2.87 | 3.54 | 3.59 | 3.22 | 2.91 | 2.63 | 3.89 | 2.69 | 0.00 | 3.43 |
| 12 | 3.28 | 3.55 | 3.81 | 4.72 | 3.34 | 3.22 | 3.84 | 3.67 | 3.97 | 3.38 | 3.43 | 0.00 |
The table contains RMSD values (calculated on the Cα atoms) between the predicted models.
Read more about the root-mean-square deviation (RMSD) measure.
| # | 1 | 2 | 3 | 4 | 5 | 6 | 7 | 8 | 9 | 10 | 11 | 12 |
| 1 | 1.00 | 0.59 | 0.56 | 0.50 | 0.57 | 0.61 | 0.57 | 0.56 | 0.69 | 0.63 | 0.62 | 0.63 |
| 2 | 0.59 | 1.00 | 0.56 | 0.50 | 0.63 | 0.67 | 0.56 | 0.57 | 0.56 | 0.56 | 0.56 | 0.60 |
| 3 | 0.56 | 0.56 | 1.00 | 0.54 | 0.53 | 0.57 | 0.59 | 0.67 | 0.58 | 0.61 | 0.67 | 0.56 |
| 4 | 0.50 | 0.50 | 0.54 | 1.00 | 0.55 | 0.53 | 0.60 | 0.57 | 0.50 | 0.52 | 0.58 | 0.50 |
| 5 | 0.57 | 0.63 | 0.53 | 0.55 | 1.00 | 0.68 | 0.56 | 0.57 | 0.56 | 0.56 | 0.58 | 0.61 |
| 6 | 0.61 | 0.67 | 0.57 | 0.53 | 0.68 | 1.00 | 0.56 | 0.59 | 0.58 | 0.59 | 0.60 | 0.62 |
| 7 | 0.57 | 0.56 | 0.59 | 0.60 | 0.56 | 0.56 | 1.00 | 0.63 | 0.54 | 0.59 | 0.63 | 0.54 |
| 8 | 0.56 | 0.57 | 0.67 | 0.57 | 0.57 | 0.59 | 0.63 | 1.00 | 0.57 | 0.61 | 0.70 | 0.54 |
| 9 | 0.69 | 0.56 | 0.58 | 0.50 | 0.56 | 0.58 | 0.54 | 0.57 | 1.00 | 0.65 | 0.63 | 0.60 |
| 10 | 0.63 | 0.56 | 0.61 | 0.52 | 0.56 | 0.59 | 0.59 | 0.61 | 0.65 | 1.00 | 0.66 | 0.58 |
| 11 | 0.62 | 0.56 | 0.67 | 0.58 | 0.58 | 0.60 | 0.63 | 0.70 | 0.63 | 0.66 | 1.00 | 0.58 |
| 12 | 0.63 | 0.60 | 0.56 | 0.50 | 0.61 | 0.62 | 0.54 | 0.54 | 0.60 | 0.58 | 0.58 | 1.00 |
The table contains GDT_TS values (calculated on the Cα atoms) between the predicted models.
Read more about the global distance test (GDT, also written as GDT_TS to represent "total score") measure.
© Laboratory of Theory of Biopolymers, Faculty of Chemistry, University of Warsaw 2013