| Chain sequence(s) |
A: MRSGSHHHHHHRSDITSLYKKAGSAAAPFTMENLYFQSYQGNSDCYFGNGSAYRGTHSLTESGASCLPWNSMILIGKVYTAQNPSAQALGLGKHNYCRNPDGDAKPWCHVLKNRRLTWEYCDVPSCSTCGLRQYSQPQFRIKGGLFADIASHPWQAAIFAKHRRSPGERFLCGGILISSCWILSAAHCFQERFPPHHLTVILGRTYRVVPGEEEQKFEVEKYIVHKEFDDDTYDNDIALLQLKSDSSRCAQESSVVRTVCLPPADLQLPDWTECELSGYGKHEALSPFYSERLKEAHVRLYPSSRCTSQHLLNRTVTDNMLCAGDTRSGGPQANLHDACQGDSGGPLVCLNDGRMTLVGIISWGLGCGQKDVPGVYTKVTNYLDWIRDNMRP |
| Distance of aggregation | 10 Å |
| Dynamic mode | Yes |
The table below lists A3D score for protein residues. Residues with A3D score > 0.0000 are marked by yellow rows.
| residue index | residue name | chain | Aggrescan3D score | mutation |
|---|---|---|---|---|
| residue index | residue name | chain | Aggrescan3D score | |
| 1 | M | A | -0.0731 | |
| 2 | R | A | -1.8700 | |
| 3 | S | A | -1.0180 | |
| 4 | G | A | -0.7333 | |
| 5 | S | A | -0.9257 | |
| 6 | H | A | -1.2045 | |
| 7 | H | A | -1.3132 | |
| 8 | H | A | -2.0180 | |
| 9 | H | A | -2.0872 | |
| 10 | H | A | -2.1319 | |
| 11 | H | A | -2.6410 | |
| 12 | R | A | -2.7973 | |
| 13 | S | A | -1.8290 | |
| 14 | D | A | -1.9011 | |
| 15 | I | A | 0.9356 | |
| 16 | T | A | 0.8889 | |
| 17 | S | A | 1.0265 | |
| 18 | L | A | 2.2214 | |
| 19 | Y | A | 1.0611 | |
| 20 | K | A | -0.8656 | |
| 21 | K | A | -1.8303 | |
| 22 | A | A | -0.8922 | |
| 23 | G | A | -0.8046 | |
| 24 | S | A | -0.3657 | |
| 25 | A | A | 0.3854 | |
| 26 | A | A | 1.0315 | |
| 27 | A | A | 0.4197 | |
| 28 | P | A | 0.5794 | |
| 29 | F | A | 1.4244 | |
| 30 | T | A | 0.9618 | |
| 31 | M | A | 0.8422 | |
| 32 | E | A | -0.9851 | |
| 33 | N | A | -1.0124 | |
| 34 | L | A | 1.4367 | |
| 35 | Y | A | 2.2342 | |
| 36 | F | A | 2.5429 | |
| 37 | Q | A | 0.7223 | |
| 38 | S | A | 0.1843 | |
| 39 | Y | A | 0.0000 | |
| 40 | Q | A | -1.1567 | |
| 41 | G | A | -1.0442 | |
| 42 | N | A | -0.7062 | |
| 43 | S | A | 0.0000 | |
| 44 | D | A | -0.2154 | |
| 45 | C | A | 0.5033 | |
| 46 | Y | A | 0.3012 | |
| 47 | F | A | -0.1666 | |
| 48 | G | A | -0.7384 | |
| 49 | N | A | -1.6598 | |
| 50 | G | A | -1.4526 | |
| 51 | S | A | -0.8912 | |
| 52 | A | A | -0.5263 | |
| 53 | Y | A | 0.0000 | |
| 54 | R | A | -0.8810 | |
| 55 | G | A | 0.0000 | |
| 56 | T | A | -1.1245 | |
| 57 | H | A | -0.5091 | |
| 58 | S | A | 0.0000 | |
| 59 | L | A | 1.3691 | |
| 60 | T | A | 0.0000 | |
| 61 | E | A | 0.0000 | |
| 62 | S | A | -0.5129 | |
| 63 | G | A | 0.0863 | |
| 64 | A | A | 0.6702 | |
| 65 | S | A | 1.3727 | |
| 66 | C | A | 1.3744 | |
| 67 | L | A | 1.9312 | |
| 68 | P | A | 0.0000 | |
| 69 | W | A | 1.0913 | |
| 70 | N | A | 0.0000 | |
| 71 | S | A | -0.1490 | |
| 72 | M | A | 0.7181 | |
| 73 | I | A | 0.0000 | |
| 74 | L | A | 0.0000 | |
| 75 | I | A | 1.0608 | |
| 76 | G | A | 0.2718 | |
| 77 | K | A | -0.5756 | |
| 78 | V | A | 1.1012 | |
| 79 | Y | A | 0.9736 | |
| 80 | T | A | 0.0000 | |
| 81 | A | A | 0.0069 | |
| 82 | Q | A | -1.4165 | |
| 83 | N | A | -1.8232 | |
| 84 | P | A | -0.9959 | |
| 85 | S | A | -1.1104 | |
| 86 | A | A | -1.3536 | |
| 87 | Q | A | -1.4411 | |
| 88 | A | A | -0.6646 | |
| 89 | L | A | -0.7315 | |
| 90 | G | A | 0.0000 | |
| 91 | L | A | 0.0000 | |
| 92 | G | A | -1.8393 | |
| 93 | K | A | -2.2731 | |
| 94 | H | A | -1.5982 | |
| 95 | N | A | -0.5544 | |
| 96 | Y | A | 0.0000 | |
| 97 | C | A | 0.0000 | |
| 98 | R | A | 0.0000 | |
| 99 | N | A | -1.1933 | |
| 100 | P | A | 0.0000 | |
| 101 | D | A | -1.9149 | |
| 102 | G | A | -1.9192 | |
| 103 | D | A | -1.9860 | |
| 104 | A | A | 0.0000 | |
| 105 | K | A | -1.4959 | |
| 106 | P | A | 0.0000 | |
| 107 | W | A | 0.0000 | |
| 108 | C | A | 0.0000 | |
| 109 | H | A | 0.0000 | |
| 110 | V | A | 0.0071 | |
| 111 | L | A | -0.9706 | |
| 112 | K | A | -3.1365 | |
| 113 | N | A | -3.4246 | |
| 114 | R | A | -3.5429 | |
| 115 | R | A | -3.5867 | |
| 116 | L | A | -1.4718 | |
| 117 | T | A | -0.8140 | |
| 118 | W | A | 0.1003 | |
| 119 | E | A | 0.0000 | |
| 120 | Y | A | -0.0551 | |
| 121 | C | A | 0.0000 | |
| 122 | D | A | -0.4391 | |
| 123 | V | A | 0.1616 | |
| 124 | P | A | -0.1107 | |
| 125 | S | A | 0.0455 | |
| 126 | C | A | 0.4226 | |
| 127 | S | A | -0.0049 | |
| 128 | T | A | -0.0223 | |
| 129 | C | A | 0.3624 | |
| 130 | G | A | 0.0000 | |
| 131 | L | A | -0.0461 | |
| 132 | R | A | 0.0000 | |
| 133 | Q | A | -0.9913 | |
| 134 | Y | A | -0.8906 | |
| 135 | S | A | -0.9653 | |
| 136 | Q | A | -1.2649 | |
| 137 | P | A | -0.7288 | |
| 138 | Q | A | 0.0000 | |
| 139 | F | A | 0.0000 | |
| 140 | R | A | 0.0000 | |
| 141 | I | A | 0.0000 | |
| 142 | K | A | 0.0000 | |
| 143 | G | A | -0.3639 | |
| 144 | G | A | 0.2358 | |
| 145 | L | A | 1.4041 | |
| 146 | F | A | 1.6548 | |
| 147 | A | A | 0.0000 | |
| 148 | D | A | -1.4660 | |
| 149 | I | A | -0.6718 | |
| 150 | A | A | -0.6573 | |
| 151 | S | A | -0.9885 | |
| 152 | H | A | 0.0000 | |
| 153 | P | A | 0.0000 | |
| 154 | W | A | 0.0000 | |
| 155 | Q | A | -0.0672 | |
| 156 | A | A | 0.0000 | |
| 157 | A | A | 0.0000 | |
| 158 | I | A | 0.0000 | |
| 159 | F | A | 0.0000 | |
| 160 | A | A | 0.0000 | |
| 161 | K | A | -3.5319 | |
| 162 | H | A | -3.2742 | |
| 163 | R | A | -4.0541 | |
| 164 | R | A | -3.4518 | |
| 165 | S | A | -2.7272 | |
| 166 | P | A | -2.7347 | |
| 167 | G | A | 0.0000 | |
| 168 | E | A | -3.5791 | |
| 169 | R | A | -3.0835 | |
| 170 | F | A | -1.1477 | |
| 171 | L | A | -0.5677 | |
| 172 | C | A | 0.0000 | |
| 173 | G | A | 0.0000 | |
| 174 | G | A | 0.0000 | |
| 175 | I | A | 0.0000 | |
| 176 | L | A | 0.0000 | |
| 177 | I | A | 0.0000 | |
| 178 | S | A | -0.1797 | |
| 179 | S | A | -0.6891 | |
| 180 | C | A | 0.0000 | |
| 181 | W | A | -0.3553 | |
| 182 | I | A | 0.0000 | |
| 183 | L | A | 0.0000 | |
| 184 | S | A | 0.0000 | |
| 185 | A | A | 0.0000 | |
| 186 | A | A | 0.0000 | |
| 187 | H | A | -1.8230 | |
| 188 | C | A | 0.0000 | |
| 189 | F | A | 0.0000 | |
| 190 | Q | A | -2.0278 | |
| 191 | E | A | -1.9865 | |
| 192 | R | A | -1.3875 | |
| 193 | F | A | -1.1033 | |
| 194 | P | A | -1.2426 | |
| 195 | P | A | -1.5685 | |
| 196 | H | A | -2.2856 | |
| 197 | H | A | -2.3398 | |
| 198 | L | A | 0.0000 | |
| 199 | T | A | 0.0000 | |
| 200 | V | A | 0.0000 | |
| 201 | I | A | -1.1983 | |
| 202 | L | A | 0.0000 | |
| 203 | G | A | 0.0000 | |
| 204 | R | A | 0.0000 | |
| 205 | T | A | -0.9098 | |
| 206 | Y | A | 0.0460 | |
| 207 | R | A | 0.4801 | |
| 208 | V | A | 2.0943 | |
| 209 | V | A | 1.7719 | |
| 210 | P | A | -0.0573 | |
| 211 | G | A | -0.6143 | |
| 212 | E | A | -2.3391 | |
| 213 | E | A | -2.7311 | |
| 214 | E | A | -2.4721 | |
| 215 | Q | A | -2.2917 | |
| 216 | K | A | -2.5379 | |
| 217 | F | A | 0.0000 | |
| 218 | E | A | -2.9090 | |
| 219 | V | A | 0.0000 | |
| 220 | E | A | -2.5105 | |
| 221 | K | A | -1.9127 | |
| 222 | Y | A | -0.2430 | |
| 223 | I | A | 0.8060 | |
| 224 | V | A | 0.1649 | |
| 225 | H | A | -1.2360 | |
| 226 | K | A | -2.6492 | |
| 227 | E | A | -3.0465 | |
| 228 | F | A | -2.3697 | |
| 229 | D | A | -3.1844 | |
| 230 | D | A | -2.6284 | |
| 231 | D | A | -2.9770 | |
| 232 | T | A | -2.3164 | |
| 233 | Y | A | -1.6832 | |
| 234 | D | A | -1.8882 | |
| 235 | N | A | -1.4836 | |
| 236 | D | A | 0.0000 | |
| 237 | I | A | 0.0000 | |
| 238 | A | A | 0.0000 | |
| 239 | L | A | 0.0000 | |
| 240 | L | A | 0.0000 | |
| 241 | Q | A | -1.2095 | |
| 242 | L | A | 0.0000 | |
| 243 | K | A | -2.5243 | |
| 244 | S | A | -2.4021 | |
| 245 | D | A | -2.3474 | |
| 246 | S | A | -1.3886 | |
| 247 | S | A | -1.4121 | |
| 248 | R | A | -1.6665 | |
| 249 | C | A | 0.0000 | |
| 250 | A | A | -1.1189 | |
| 251 | Q | A | -1.7280 | |
| 252 | E | A | -1.5999 | |
| 253 | S | A | -0.9643 | |
| 254 | S | A | -0.4317 | |
| 255 | V | A | -0.8352 | |
| 256 | V | A | 0.0000 | |
| 257 | R | A | -0.6125 | |
| 258 | T | A | 0.0000 | |
| 259 | V | A | 0.0000 | |
| 260 | C | A | 0.2217 | |
| 261 | L | A | 0.0000 | |
| 262 | P | A | -0.7320 | |
| 263 | P | A | -1.2612 | |
| 264 | A | A | -1.3324 | |
| 265 | D | A | -1.6668 | |
| 266 | L | A | -0.5740 | |
| 267 | Q | A | -0.9841 | |
| 268 | L | A | -0.0701 | |
| 269 | P | A | -0.7774 | |
| 270 | D | A | -1.7642 | |
| 271 | W | A | 0.0000 | |
| 272 | T | A | -0.9107 | |
| 273 | E | A | -1.1793 | |
| 274 | C | A | 0.0000 | |
| 275 | E | A | 0.0000 | |
| 276 | L | A | 0.0000 | |
| 277 | S | A | 0.0000 | |
| 278 | G | A | 0.0000 | |
| 279 | Y | A | 0.0000 | |
| 280 | G | A | 0.0000 | |
| 281 | K | A | -0.5062 | |
| 282 | H | A | 0.0000 | |
| 283 | E | A | 0.3658 | |
| 284 | A | A | 0.7293 | |
| 285 | L | A | 1.3702 | |
| 286 | S | A | 0.9216 | |
| 287 | P | A | 0.7370 | |
| 288 | F | A | 1.6085 | |
| 289 | Y | A | 1.3519 | |
| 290 | S | A | 0.5997 | |
| 291 | E | A | 0.0000 | |
| 292 | R | A | 0.3915 | |
| 293 | L | A | 0.0000 | |
| 294 | K | A | 0.0000 | |
| 295 | E | A | 0.0000 | |
| 296 | A | A | 0.0000 | |
| 297 | H | A | -1.0524 | |
| 298 | V | A | 0.0000 | |
| 299 | R | A | -2.0871 | |
| 300 | L | A | 0.0000 | |
| 301 | Y | A | -0.9729 | |
| 302 | P | A | -0.7100 | |
| 303 | S | A | -0.7392 | |
| 304 | S | A | -1.1441 | |
| 305 | R | A | -2.1221 | |
| 306 | C | A | 0.0000 | |
| 307 | T | A | -1.2145 | |
| 308 | S | A | -1.3564 | |
| 309 | Q | A | -1.5492 | |
| 310 | H | A | -1.2564 | |
| 311 | L | A | 0.0000 | |
| 312 | L | A | -0.2863 | |
| 313 | N | A | -1.9221 | |
| 314 | R | A | -2.2628 | |
| 315 | T | A | -1.4701 | |
| 316 | V | A | -0.6781 | |
| 317 | T | A | -0.4784 | |
| 318 | D | A | -0.4257 | |
| 319 | N | A | -0.3543 | |
| 320 | M | A | 0.3842 | |
| 321 | L | A | 0.0000 | |
| 322 | C | A | 0.0000 | |
| 323 | A | A | 0.0000 | |
| 324 | G | A | -1.4293 | |
| 325 | D | A | 0.0000 | |
| 326 | T | A | -2.3348 | |
| 327 | R | A | -3.0427 | |
| 328 | S | A | -2.2983 | |
| 329 | G | A | -1.6654 | |
| 330 | G | A | -1.4668 | |
| 331 | P | A | -1.1869 | |
| 332 | Q | A | -1.7088 | |
| 333 | A | A | -1.3933 | |
| 334 | N | A | -1.8335 | |
| 335 | L | A | -1.3877 | |
| 336 | H | A | -0.7681 | |
| 337 | D | A | 0.0000 | |
| 338 | A | A | 0.0000 | |
| 339 | C | A | -0.7276 | |
| 340 | Q | A | -1.0796 | |
| 341 | G | A | -0.6404 | |
| 342 | D | A | 0.0000 | |
| 343 | S | A | -0.4577 | |
| 344 | G | A | 0.0000 | |
| 345 | G | A | 0.0000 | |
| 346 | P | A | 0.0000 | |
| 347 | L | A | 0.0000 | |
| 348 | V | A | 0.0000 | |
| 349 | C | A | 0.1359 | |
| 350 | L | A | 0.1221 | |
| 351 | N | A | -1.4608 | |
| 352 | D | A | -2.0507 | |
| 353 | G | A | -1.2167 | |
| 354 | R | A | -1.1244 | |
| 355 | M | A | 0.0000 | |
| 356 | T | A | 0.0512 | |
| 357 | L | A | 0.0000 | |
| 358 | V | A | 0.0000 | |
| 359 | G | A | 0.0000 | |
| 360 | I | A | 0.0000 | |
| 361 | I | A | 0.0000 | |
| 362 | S | A | 0.0000 | |
| 363 | W | A | -0.2813 | |
| 364 | G | A | -0.6110 | |
| 365 | L | A | -0.0285 | |
| 366 | G | A | -0.3887 | |
| 367 | C | A | -0.3641 | |
| 368 | G | A | -1.2492 | |
| 369 | Q | A | -2.2644 | |
| 370 | K | A | -3.0572 | |
| 371 | D | A | -2.3379 | |
| 372 | V | A | 0.0000 | |
| 373 | P | A | 0.0000 | |
| 374 | G | A | 0.0000 | |
| 375 | V | A | 0.0000 | |
| 376 | Y | A | 0.0000 | |
| 377 | T | A | 0.0000 | |
| 378 | K | A | -0.3523 | |
| 379 | V | A | 0.0000 | |
| 380 | T | A | -0.8859 | |
| 381 | N | A | -1.1827 | |
| 382 | Y | A | 0.0000 | |
| 383 | L | A | 0.0000 | |
| 384 | D | A | -1.9498 | |
| 385 | W | A | -0.8897 | |
| 386 | I | A | 0.0000 | |
| 387 | R | A | -3.0384 | |
| 388 | D | A | -3.0978 | |
| 389 | N | A | -1.9654 | |
| 390 | M | A | -1.6892 | |
| 391 | R | A | -2.7479 | |
| 392 | P | A | -1.6679 |
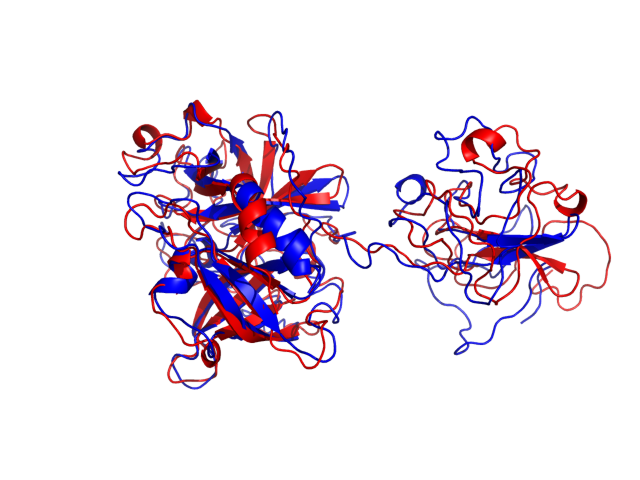
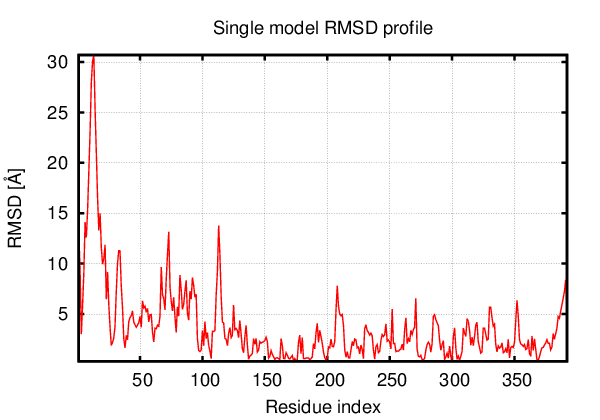
Above is the comparison between the input structure and the most aggregation prone model (predicted in the flexibility simulations, download both models in the PDB file format , download table with rmsd values ). The picture presents the most aggregation prone model (in blue) superimposed on the input structure (in red). The plot shows rmsd profile (distances between residues of the superimposed structures).
Dynamic mode uses CABS-flex simulations of protein structure fluctuations. The structure fluctuations may have impact on the size and extent of aggregation "hot-spots" on the protein surface. The dynamic mode uses the following pipeline: (1) based on the input structure, CABS-flex predicts a set of different models reflecting protein dynamics in solution; (2) for each of these models A3D score is calculated; (3) finally, the most aggregation prone model (the model with the highest A3D score, -0.6348 in this case) is selected and presented in A3D results.